multiBigwigSummary¶
Given two or more bigWig files, multiBigwigSummary computes the average scores for each of the files in every genomic region. This analysis is performed for the entire genome by running the program in ‘bins’ mode, or for certain user selected regions in ‘BED-file’ mode. Most commonly, the output of multiBigwigSummary is used by other tools such as ‘plotCorrelation’ or ‘plotPCA’ for visualization and diagnostic purposes.
A detailed sub-commands help is available by typing:
multiBigwigSummary bins -h
multiBigwigSummary BED-file -h
usage: multiBigwigSummary [-h] [--version] ...
- optional arguments
--version show program’s version number and exit - commands
Undocumented
Possible choices: bins, BED-file
- Sub-commands:
- bins
The average score is based on equally sized bins (10 kilobases by default), which consecutively cover the entire genome. The only exception is the last bin of a chromosome, which is often smaller. The output of this mode is commonly used to assess the overall similarity of different bigWig files.
usage: multiBigwigSummary -b file1.bw file2.bw -out results.npz
- Required arguments
--bwfiles, -b List of bigWig files, separated by spaces. --outFileName, -out File name to save the compressed matrix file (npz format)needed by the “plotHeatmap” and “plotProfile” tools. - Optional arguments
--labels, -l User defined labels instead of default labels from file names. Multiple labels have to be separated by spaces, e.g., –labels sample1 sample2 sample3 --chromosomesToSkip List of chromosomes that you do not want to be included for the correlation. Useful to remove “random” or “extra” chr. --binSize=10000, -bs=10000 Size (in bases) of the windows sampled from the genome. --distanceBetweenBins=0, -n=0 By default, multiBigwigSummary considers adjacent bins of the specified –binSize. However, to reduce the computation time, a larger distance between bins can be given. Larger distances results in fewer considered bins. --version show program’s version number and exit --region, -r Region of the genome to limit the operation to - this is useful when testing parameters to reduce the computing time. The format is chr:start:end, for example –region chr10 or –region chr10:456700:891000. --numberOfProcessors=max/2, -p=max/2 Number of processors to use. Type “max/2” to use half the maximum number of processors or “max” to use all available processors. --verbose=False, -v=False Set to see processing messages. - Output optional options
--outRawCounts Save average scores per region for each bigWig file to a single tab-delimited file.
- BED-file
The user provides a BED file that contains all regions that should be considered for the analysis. A common use is to compare scores (e.g. ChIP-seq scores) between different samples over a set of pre-defined peak regions.
usage: multiBigwigSummary -b file1.bw file2.bw -out results.npz --BED selection.bed
- Required arguments
--bwfiles, -b List of bigWig files, separated by spaces. --outFileName, -out File name to save the compressed matrix file (npz format)needed by the “plotHeatmap” and “plotProfile” tools. --BED Limits the correlation analysis to the regions specified in this file. - Optional arguments
--labels, -l User defined labels instead of default labels from file names. Multiple labels have to be separated by spaces, e.g., –labels sample1 sample2 sample3 --chromosomesToSkip List of chromosomes that you do not want to be included for the correlation. Useful to remove “random” or “extra” chr. --version show program’s version number and exit --region, -r Region of the genome to limit the operation to - this is useful when testing parameters to reduce the computing time. The format is chr:start:end, for example –region chr10 or –region chr10:456700:891000. --numberOfProcessors=max/2, -p=max/2 Number of processors to use. Type “max/2” to use half the maximum number of processors or “max” to use all available processors. --verbose=False, -v=False Set to see processing messages. - Output optional options
--outRawCounts Save average scores per region for each bigWig file to a single tab-delimited file.
- example usages:
- multiBigwigSummary bins -b file1.bw file2.bw -out results.npz
multiBigwigSummary BED-file -b file1.bw file2.bw -out results.npz –BED selection.bed
Example¶
In the following example, the average values for our test ENCODE ChIP-Seq datasets are computed for consecutive genome bins (default size: 10kb) by using the bins mode.
$ deepTools2.0/bin/multiBigwigSummary bins \
-b testFiles/H3K4Me1.bigWig testFiles/H3K4Me3.bigWig testFiles/H3K27Me3.bigWig testFiles/Input.bigWig \
--labels H3K4me1 H3K4me3 H3K27me3 input \
-out scores_per_bin.npz --outRawCounts scores_per_bin.tab
$ head scores_per_bin.tab
#'chr' 'start' 'end' 'H3K4me1' 'H3K4me3' 'H3K27me3' 'input'
19 0 10000 0.0 0.0 0.0 0.0
19 10000 20000 0.0 0.0 0.0 0.0
19 20000 30000 0.0 0.0 0.0 0.0
19 30000 40000 0.0 0.0 0.0 0.0
19 40000 50000 0.0 0.0 0.0 0.0
19 50000 60000 0.0221538461538 0.0 0.00482142857143 0.0522717391304
19 60000 70000 4.27391282051 1.625 0.634116071429 1.29124347826
19 70000 80000 13.0891675214 24.65 1.8180625 2.80073695652
19 80000 90000 1.74591965812 0.29 4.35576785714 0.92987826087
To compute the average values for a set of genes, use the BED-file mode.
$ deepTools2.0/bin/multiBigwigSummary BED-file \
--bwfiles testFiles/*bigWig \
--BED testFiles/genes.bed \
--labels H3K27me3 H3K4me1 H3K4me3 HeK9me3 input \
-out scores_per_transcript.npz --outRawCounts scores_per_transcript.tab
$ head scores_per_transcript.tab
#'chr' 'start' 'end' 'H3K27me3' 'H3K4me1' 'H3K4me3' 'HeK9me3' 'input'
19 60104 70951 0.663422099099 4.37103606574 14.9609108509 0.596631607217 1.34274297191
19 60950 70966 0.714223982699 4.54650763906 16.2336261981 0.62173674295 1.41719308888
19 62114 70944 0.747578769617 4.84009060023 18.2951302378 0.648723472352 1.51324474371
19 63820 70951 0.781816722009 5.30500631048 22.5579862572 0.682862029229 1.55490104062
19 65057 66382 0.528301886792 5.45886792453 0.523018867925 0.555471698113 1.97056603774
19 65821 66416 0.411764705882 3.0 0.636974789916 0.168067226891 1.67226890756
19 65821 70945 0.844600775761 4.79176424668 31.1346604215 0.693073728066 1.47911787666
19 66319 66492 0.774566473988 1.59537572254 0.0 0.0 0.578034682081
19 66345 71535 0.877430197151 5.49036608863 43.978805395 0.746026011561 1.43545279383
The default output of multiBamCoverage (a compressed numpy array: *.npz) can be visualized using plotCorrelation or plotPCA.
The optional output (--outRawCounts) is a simple tab-delimited file that can be used with any other program. The first three columns define the region of the genome for which the reads were summarized.
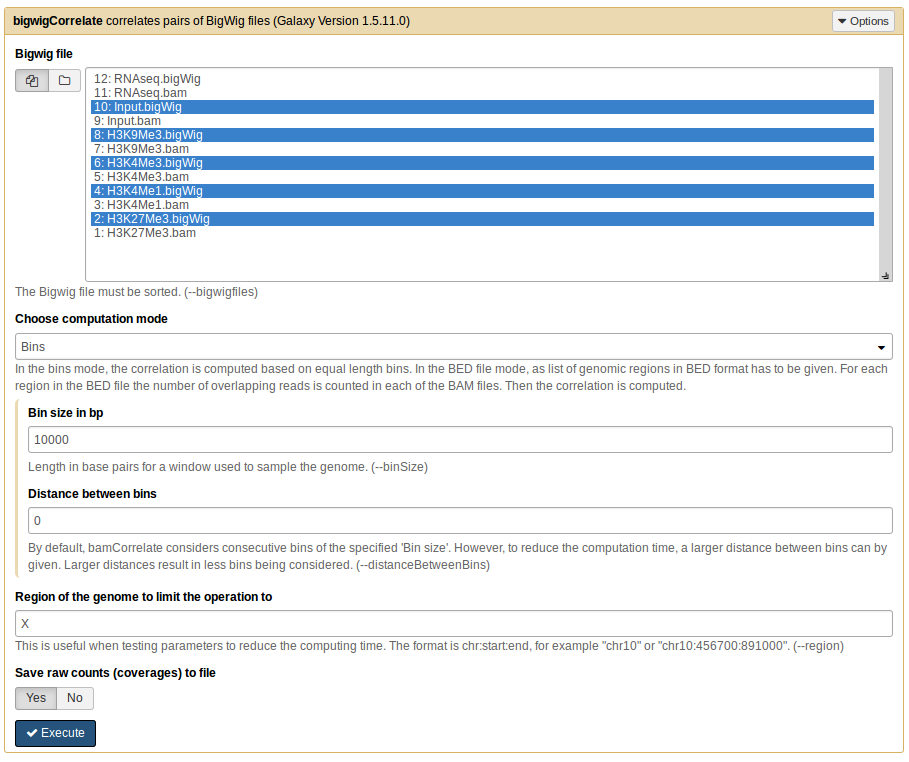
multiBigwigSummary in Galaxy¶
Below is the screenshot showing how to use multiBigwigSummary on the deeptools galaxy.