plotPCA¶
Tool for generating a principal component analysis (PCA) plot from multiBamSummary or multiBigwigSummary output.
Detailed help:
plotPCA -h
usage: plotPCA [-h] --corData FILE --plotFile FILE
[--labels sample1 sample2 [sample1 sample2 ...]]
[--plotTitle PLOTTITLE] [--plotFileFormat FILETYPE]
[--outFileNameData OUTFILENAMEDATA] [--rowCenter] [--version]
Required arguments¶
| –corData, -in | Coverage file (generated by multiBamSummary or multiBigwigSummary) |
| –plotFile, -o | File name to save the plot to. The extension determines the file format. For example: pca.pdf will save the PCA plot in PDF format. The available options are: .png, .eps, .pdf and .svg. |
Optional arguments¶
| –labels, -l | User defined labels instead of default labels from file names. Multiple labels have to be separated by spaces, e.g. –labels sample1 sample2 sample3 |
| –plotTitle, -T | |
| Title of the plot, to be printed on top of the generated image. Leave blank for no title. | |
| –plotFileFormat | |
Possible choices: png, pdf, svg, eps Image format type. If given, this option overrides the image format based on the plotFile ending. The available options are: png, eps, pdf and svg. | |
| –outFileNameData | |
| File name to save the data underlying data for the average profile, e.g., myProfile.tab. | |
| –rowCenter | When specified, each row (bin, gene, etc.) in the matrix is centered at 0 before the PCA is computed. This is useful only if you have a strong bin/gene/etc. correlation and the resulting principal component has samples stacked vertically. |
| –version | show program’s version number and exit |
example usages: plotPCA -in coverages.npz -o pca.png
Background¶
Principal component analysis (PCA) can be used, for example, to determine whether samples display greater variability between experimental conditions than between replicates of the same treatment. PCA is also useful to identify unexpected patterns, such as those caused by batch effects or outliers. Principal components represent the directions along which the variation in the data is maximal, so that the information (e.g., read coverage values) from thousands of regions can be represented by just a few dimensions.
Note
PCA is not designed to identify unknown groupings or clustering and given an unexpected result, it is up to the researcher to determine the experimental or technical reason underlying the principal components.
Usage example¶
plotPCA needs the compressed numpy array output from either multiBamSummary or multiBigwigSummary
$ deepTools2.0/bin/plotPCA -in readCounts.npz \
-o PCA_readCounts.png \
-T "PCA of read counts"
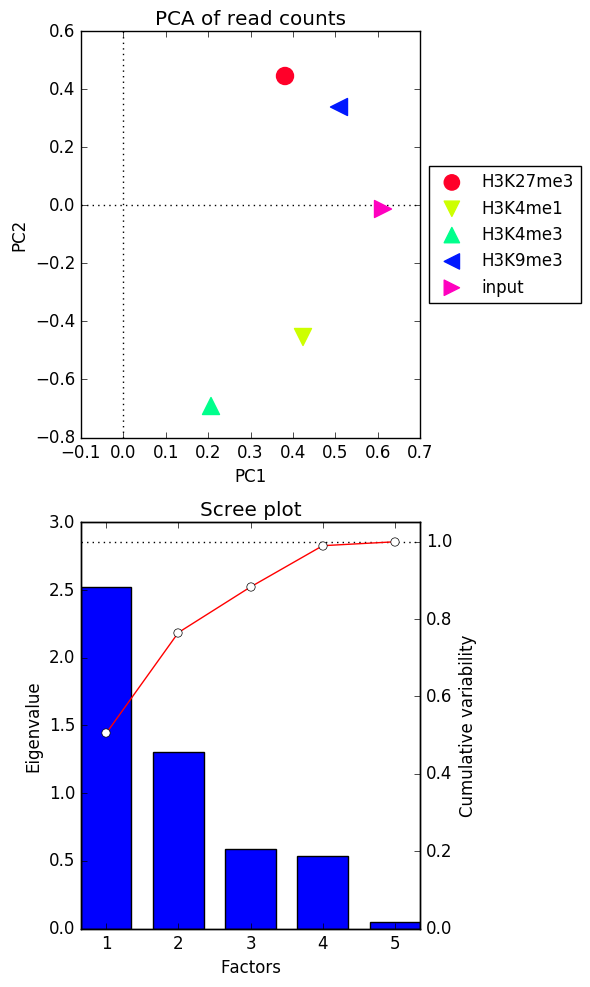
After perfoming the PCA on the values supplied as the input, plotPCA will sort the principal components according to the amount of variability of the data that they explain. Based on this, you will obtain two plots:
- the eigenvalues of the top two principal components
- the Scree plot for the top five principal components where the bars represent the amount of variability explained by the individual factors and the red line traces the amount of variability is explained by the individual components in a cumulative manner
| deepTools Galaxy. | code @ github. |