bamCompare
bamCompare can be used to generate a bigWig or bedGraph file based on two BAM files that are compared to each other while being simultaneously normalized for sequencing depth.
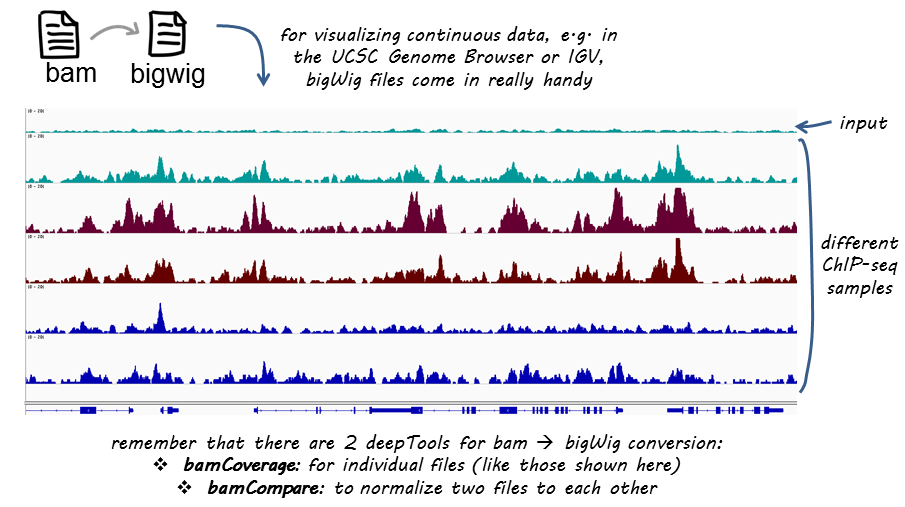
If you are not familiar with BAM, bedGraph and bigWig formats, you can read up on that in our Glossary of NGS terms
The basic algorithm works proceeds in two steps:
Per-sample scaling / depth Normalization:
If scaling is used (using the SES or read counts method), appropriate scaling factors are determined to account for sequencing depth differences.
Optionally scaling can be turned off and individual samples normalized using the RPKM, BPM or CPM methods (or no normalization at all)
A per-bin calculation is performed after accounting for scaling:
The genome is transversed and the log2 ratio/ratio/difference/etc. for each bin of fixed width is computed.
This tool compares two BAM files based on the number of mapped reads. To compare the BAM files, the genome is partitioned into bins of equal size, then the number of reads found in each bin is counted per file, and finally a summary value is reported. This value can be the ratio of the number of reads per bin, the log2 of the ratio, or the difference. This tool can normalize the number of reads in each BAM file using the SES method proposed by Diaz et al. (2012) “Normalization, bias correction, and peak calling for ChIP-seq”. Statistical Applications in Genetics and Molecular Biology, 11(3). Normalization based on read counts is also available. The output is either a bedgraph or bigWig file containing the bin location and the resulting comparison value. Note that each end in a pair (for paired-end reads) is treated independently. If this is undesirable, then use the –samFlagInclude or –samFlagExclude options.
usage: bamCompare -b1 treatment.bam -b2 control.bam -o log2ratio.bw
help: bamCompare -h / bamCompare --help
Required arguments
- --bamfile1, -b1
Sorted BAM file 1. Usually the BAM file for the treatment.
- --bamfile2, -b2
Sorted BAM file 2. Usually the BAM file for the control.
Output
- --outFileName, -o
Output file name.
- --outFileFormat, -of
Possible choices: bigwig, bedgraph
Output file type. Either “bigwig” or “bedgraph”.
Optional arguments
- --scaleFactorsMethod
Possible choices: readCount, SES, None
Method to use to scale the samples. If a method is specified, then it will be used to compensate for sequencing depth differences between the samples. As an alternative, this can be set to None and an option from –normalizeUsing <method> can be used. (Default: “readCount”)
- --sampleLength, -l
Only relevant when SES is chosen for the scaleFactorsMethod. To compute the SES, specify the length (in bases) of the regions (see –numberOfSamples) that will be randomly sampled to calculate the scaling factors. If you do not have a good sequencing depth for your samples consider increasing the sampling regions’ size to minimize the probability that zero-coverage regions are used. (Default: 1000)
- --numberOfSamples, -n
Only relevant when SES is chosen for the scaleFactorsMethod. Number of samplings taken from the genome to compute the scaling factors. (Default: 100000.0)
- --scaleFactors
Set this parameter manually to avoid the computation of scaleFactors. The format is scaleFactor1:scaleFactor2.For example, –scaleFactor 0.7:1 will cause the first BAM file tobe multiplied by 0.7, while not scaling the second BAM file (multiplication with 1).
- --operation
Possible choices: log2, ratio, subtract, add, mean, reciprocal_ratio, first, second
The default is to output the log2 ratio of the two samples. The reciprocal ratio returns the the negative of the inverse of the ratio if the ratio is less than 0. The resulting values are interpreted as negative fold changes. Instead of performing a computation using both files, the scaled signal can alternatively be output for the first or second file using the ‘–operation first’ or ‘–operation second’. (Default: “log2”)
- --pseudocount
A small number to avoid x/0. Only useful together with –operation log2 or –operation ratio. You can specify different values as pseudocounts for the numerator and the denominator by providing two values (the first value is used as the numerator pseudocount and the second the denominator pseudocount). (Default: [1])
- --skipZeroOverZero
Skip bins where BOTH BAM files lack coverage. This is determined BEFORE any applicable pseudocount is added.
- --version
show program’s version number and exit
- --binSize, -bs
Size of the bins, in bases, for the output of the bigwig/bedgraph file. (Default: 50)
- --region, -r
Region of the genome to limit the operation to - this is useful when testing parameters to reduce the computing time. The format is chr:start:end, for example –region chr10 or –region chr10:456700:891000.
- --blackListFileName, -bl
A BED or GTF file containing regions that should be excluded from all analyses. Currently this works by rejecting genomic chunks that happen to overlap an entry. Consequently, for BAM files, if a read partially overlaps a blacklisted region or a fragment spans over it, then the read/fragment might still be considered. Please note that you should adjust the effective genome size, if relevant.
- --numberOfProcessors, -p
Number of processors to use. Type “max/2” to use half the maximum number of processors or “max” to use all available processors. (Default: 1)
- --verbose, -v
Set to see processing messages.
Read coverage normalization options
- --effectiveGenomeSize
The effective genome size is the portion of the genome that is mappable. Large fractions of the genome are stretches of NNNN that should be discarded. Also, if repetitive regions were not included in the mapping of reads, the effective genome size needs to be adjusted accordingly. A table of values is available here: http://deeptools.readthedocs.io/en/latest/content/feature/effectiveGenomeSize.html .
- --normalizeUsing
Possible choices: RPKM, CPM, BPM, RPGC, None
Use one of the entered methods to normalize the number of reads per bin. By default, no normalization is performed. RPKM = Reads Per Kilobase per Million mapped reads; CPM = Counts Per Million mapped reads, same as CPM in RNA-seq; BPM = Bins Per Million mapped reads, same as TPM in RNA-seq; RPGC = reads per genomic content (1x normalization); Mapped reads are considered after blacklist filtering (if applied). RPKM (per bin) = number of reads per bin / (number of mapped reads (in millions) * bin length (kb)). CPM (per bin) = number of reads per bin / number of mapped reads (in millions). BPM (per bin) = number of reads per bin / sum of all reads per bin (in millions). RPGC (per bin) = number of reads per bin / scaling factor for 1x average coverage. None = the default and equivalent to not setting this option at all. This scaling factor, in turn, is determined from the sequencing depth: (total number of mapped reads * fragment length) / effective genome size. The scaling factor used is the inverse of the sequencing depth computed for the sample to match the 1x coverage. This option requires –effectiveGenomeSize. Each read is considered independently, if you want to only count one mate from a pair in paired-end data, then use the –samFlagInclude/–samFlagExclude options. (Default: None)
- --exactScaling
Instead of computing scaling factors based on a sampling of the reads, process all of the reads to determine the exact number that will be used in the output. This requires significantly more time to compute, but will produce more accurate scaling factors in cases where alignments that are being filtered are rare and lumped together. In other words, this is only needed when region-based sampling is expected to produce incorrect results.
- --ignoreForNormalization, -ignore
A list of space-delimited chromosome names containing those chromosomes that should be excluded for computing the normalization. This is useful when considering samples with unequal coverage across chromosomes, like male samples. An usage examples is –ignoreForNormalization chrX chrM.
- --skipNonCoveredRegions, --skipNAs
This parameter determines if non-covered regions (regions without overlapping reads) in a BAM file should be skipped. The default is to treat those regions as having a value of zero. The decision to skip non-covered regions depends on the interpretation of the data. Non-covered regions may represent, for example, repetitive regions that should be skipped.
- --smoothLength
The smooth length defines a window, larger than the binSize, to average the number of reads. For example, if the –binSize is set to 20 and the –smoothLength is set to 60, then, for each bin, the average of the bin and its left and right neighbors is considered. Any value smaller than –binSize will be ignored and no smoothing will be applied.
Read processing options
- --extendReads, -e
This parameter allows the extension of reads to fragment size. If set, each read is extended, without exception. NOTE: This feature is generally NOT recommended for spliced-read data, such as RNA-seq, as it would extend reads over skipped regions. Single-end: Requires a user specified value for the final fragment length. Reads that already exceed this fragment length will not be extended. Paired-end: Reads with mates are always extended to match the fragment size defined by the two read mates. Unmated reads, mate reads that map too far apart (>4x fragment length) or even map to different chromosomes are treated like single-end reads. The input of a fragment length value is optional. If no value is specified, it is estimated from the data (mean of the fragment size of all mate reads).
- --ignoreDuplicates
If set, reads that have the same orientation and start position will be considered only once. If reads are paired, the mate’s position also has to coincide to ignore a read.
- --minMappingQuality
If set, only reads that have a mapping quality score of at least this are considered.
- --centerReads
By adding this option, reads are centered with respect to the fragment length. For paired-end data, the read is centered at the fragment length defined by the two ends of the fragment. For single-end data, the given fragment length is used. This option is useful to get a sharper signal around enriched regions.
- --samFlagInclude
Include reads based on the SAM flag. For example, to get only reads that are the first mate, use a flag of 64. This is useful to count properly paired reads only once, as otherwise the second mate will be also considered for the coverage. (Default: None)
- --samFlagExclude
Exclude reads based on the SAM flag. For example, to get only reads that map to the forward strand, use –samFlagExclude 16, where 16 is the SAM flag for reads that map to the reverse strand. (Default: None)
- --minFragmentLength
The minimum fragment length needed for read/pair inclusion. This option is primarily useful in ATACseq experiments, for filtering mono- or di-nucleosome fragments. (Default: 0)
- --maxFragmentLength
The maximum fragment length needed for read/pair inclusion. (Default: 0)